By analysing the genome of a terrestrial alga, a research group including researchers at Tokyo Institute of Technology, Kazusa DNA Research Institute and RIKEN reveal the presence of genes that enable plants to cope with the extremes of life on land, such as drought, and exposure to high-intensity light and UV.
Summary
“The colonisation of land by plants was a key event in the evolution of life,” point out researchers in Japan in a recent report. The transition provided the nutrients and oxygen that allow terrestrial creatures to survive, yet so far there has been little evidence of the stages in the evolution of plants that allowed them to adapt from aquatic life to conditions on land. A collaboration of researchers led by Hiroyuki Ohta from Center For Biological Resources and Informatics/Earth-Life Science Institute, Tokyo Institute of Technology in Japan has now identified features in the genome of a simple terrestrial alga that represent key evolutionary milestones for surviving in the more severe environments found on land.
The ancestors of present land plants are widely believed to be charophytes, a division of green algae that are distributed throughout the world. Despite the potential significance in the evolution of life, so far genome sequence data for charophytes has been fragmentary.
The charophyte Klebsormidium is a very simple multicellular organism and most species that have adapted to live on land can also live in water. Ohta and his team of researchers, which included scientists from several institutions in Japan, analysed the genome of the terrestrial Klebsormidium flaccidum (Fig. 1) and compared it with other algae and land plants.
The comparison suggests that many of the genes required to survive on land were already present in the ancestor of K flaccidum. The researchers also identified genes coding for mechanisms to respond to environmental stimuli and to protect against high intensity sunlight.
The researchers conclude their report, “Our analysis provides evidence that K flaccidum has the fundamental machinery required for adaptation to survival in terrestrial environments.”
Background
Evolution for terrestrial life
Aquatic environments are significantly more stable than the conditions on land. High-intensity light and UV radiation and drought can cause stress and damage to plant cells without the evolutionary mechanisms to cope with them.
Gene family and domain comparisons
The researchers compared the sequences of 31 proteins from K flaccidum, 5 land plants, 7 charophyte algae and 9 other algae. They compared paralogues (genes belonging to a family containing two or more genes) singletons (genes that have no paralogue) and the number of gene families (meaning the number of families of paralogues and singletons).
They found that in algae the number of gene families increased in proportion to the total number of genes, whereas for land plants the number of gene families plateaus and paralogues occur more frequently (Fig. 2). Notably the genes for important processes for terrestrial life - such as cell wall biogenesis and responding to external signals - occurred as singletons in K flaccidum, whereas they had many paralogues in land plants. They also found that a high percentage of protein domains and domain combinations found in land plants- 90.7% of the protein domains and 84.3% of the domain combinations - were present in K flaccidum (Fig. 3).
Plant hormone mediated responses for dealing with drought, salinity and freezing
Land plants respond to changes in their environment through the release of hormones, which trigger response mechanisms. For example abscissic acid (ABA) is a signalling molecule for adapting to drought, salinity and freezing and auxin and cytokinin trigger a number of mechanisms key to growth. Presence of some hormones and counterparts of a part of their receptors were identified in K flaccidum, although many well-known receptors crucial for plant hormone signalling were absent suggesting that the alga had developed primitive land-plant signalling pathways.
Cyclic electron flow for dealing with high-intensity light
Several stresses, such as high-intensity light and drought activate cyclic electron flow in what is known as a stress responsive system in higher plant photosynthesis. Cyclic electron flow causes quenching processes and ATP synthesis, which help dissipate excess light energy protecting land plants from damage. The researchers identified seven genes that encode for cyclic electron flow pathways in K flaccidum.
Reference
Koichi Hori, Fumito Maruyama, Takatomo Fujisawa, Tomoaki Togashi, Nozomi Yamamoto, Mitsunori Seo, Syusei Sato, Takuji Yamada, Hiroshi Mori, Naoyuki Tajima et al. “Klebsormidium flaccidum genome reveals primary factors for plant terrestrial adaptation” Nature Communications 5, 4978 (DOI: 10.1038/ncomms4978 ).
).
Figure image and caption
Tables and figures from the article appeared in Nature Communications.
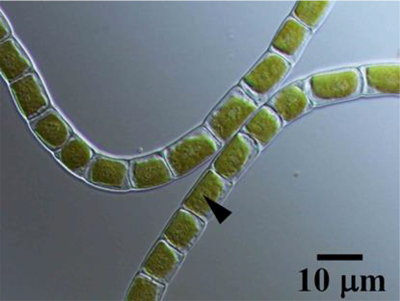
Fig. 1 Differential interference microscope image of Klebsormidium flaccidum strain NIES-2285. K. flaccidum consists of non-branching long filamentous cells. Each cell contains a large chloroplast, which is positioned against the cell wall (parietal chloroplast) and contains a pyrenoid. Arrowhead indicates a pyrenoid surrounded by a few starch grains. Scale bar = 10 μm.
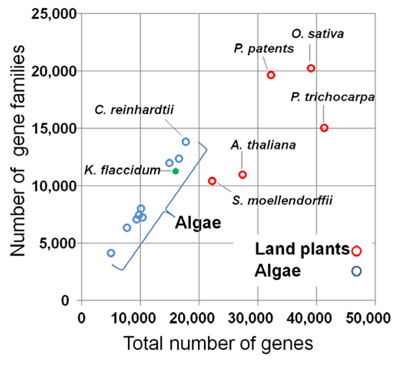
Fig. 2 Gene families in 15 species of algae and land plants. The green filled circle denotes the data point for K. flaccidum, and red and blue circles denote data points for land plants and algae, respectively.
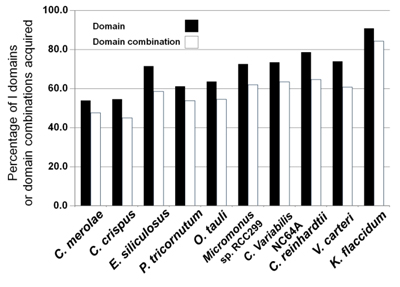
Fig. 3 Percentages of domains and domain combinations in algae that are commonly found in land plants. Acquisition in algal genomes of conserved domains (black bars) and domain combinations (white bars) commonly found in land plants. For the land plants analysed (five species), the numbers of conserved domains and domain combinations were 4,894 and 2,801, respectively.
. Any information published on this site will be valid in relation to Science Tokyo.





