A unique method to screen large-scale libraries for industrially useful bacterial strains was recently developed by Tokyo Tech researchers. The simple approach combines biosensors and microfluidics to quickly identify mutant strains that secrete large amounts of industrially useful proteins, opening the doors to more applications, like reasonably priced biopharmaceuticals.
With modern genetic engineering tools, it is now possible to modify microorganisms so that their production of industrially useful proteins—such as those used in biopharmaceuticals—is enhanced. By introducing genetic modifications into these organisms, we can use them as biological factories to produce large quantities of the desired protein. Bacteria with this enhanced ability can produce insulin, growth hormones, and enzymes. This approach of increasing microbial secretory protein expression has led to breakthroughs in medicine, industry, and agriculture.
Nonetheless, the traditional method of genetically engineering bacterial strains for high protein production is extremely time consuming. This is because it relies on introducing genetic modifications in individual strains and evaluating the effectiveness of protein production. As an alternative, researchers sometimes rely on screening of large-scale libraries for identifying strains which secrete a high amount of protein. This enables the extraction of only those strains that are the best at producing the desired protein. Unfortunately, current screening techniques rely on multiple chemical treatments and are either too slow or too complicated.
To overcome these limitations, a team of researchers has now developed a novel, high-throughput mutant strain screening method. The study was led by Associate Professor Tetsuya Kitaguchi from Tokyo Institute of Technology (Tokyo Tech), Japan, and was conducted in collaboration with Ajinomoto Co., Inc. The innovative method, which combines microfluidics and versatile biosensing to quickly identify enhanced bacterial strains that produce the highest amount of a desired protein, is reported in their study published in the journal Small on April 24, 2023.
To this end, the researchers first used a type of biosensor called Q-body to measure the amount of the desired protein produced by each strain. Q-bodies are artificial antibodies which become fluorescent on binding to their target. In this case, they were designed to bind to the desired protein, establishing a connection between fluorescence intensity and target protein production.
In addition, the team also devised a clever protocol for sorting the mutant strains based on their performance (Figure 1). Using microfluidic technology, tiny droplets of water containing individual bacteria and Q-bodies were introduced in an oil emulsion, taking advantage of oil and water’s mutual immiscibility. These tiny droplets were used as microscopic bacterial cultures and reactors.
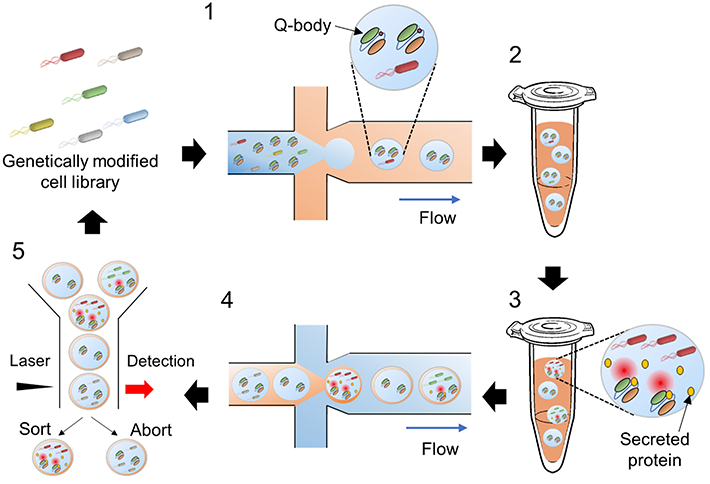
Figure 1. Overview of the screening approach
The proposed method involves using water microdroplets as tiny bacterial cultures and bioreactors and then sorting them based on their fluorescence intensity. The Q-bodies are designed to become fluorescent upon binding to a target protein. Thus, fluorescence intensity is directly related to the amount of the desired protein that each bacterial strain can produce.
After 48 hours of incubation, these oil-covered water droplets were encapsulated yet again, in a water emulsion, and sent through a flow cytometer. This device uses a laser and a detector to measure the fluorescence of each individual droplet. Following this, it employs a sorting mechanism to separate droplets with higher fluorescence intensity.
The researchers put their method to the test by screening a huge library of bacterial strains created to produce FGF9, a human cytokine, and subjected to circumstances that cause random mutations. Using this method, the team was able to identify a mutant strain that produced three times as much FGF9 compared to the control strain (Figure 2). As Dr. Kitaguchi remarks, "The entire screening process of 106 mutants was completed in approximately three days, surpassing the throughput of culture evaluation methods that use the latest automated lab instruments."
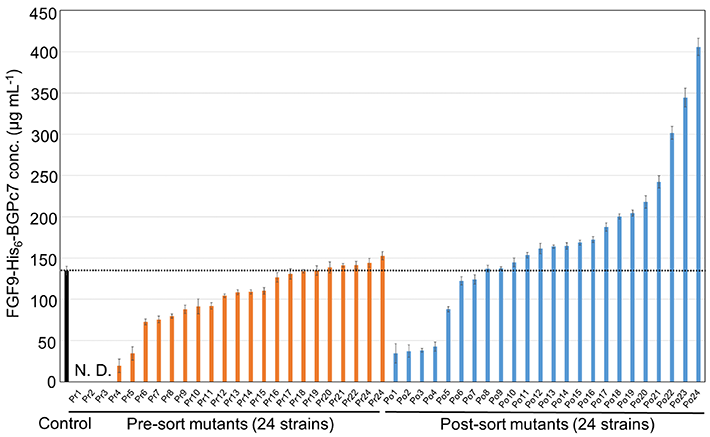
Figure 2. Finding the best performing strain among mutant bacteria
The plot shows a comparison of the amount of the desired protein produced between 24 randomly picked strains before sorting and 24 strains picked from the top 50 performers. The best-performing strain produced thrice as much of the desired protein compared to the control strain, showcasing the potential of the proposed approach.
Going ahead, the team has high expectations; they hope their proposed method will have a significant impact on the pharmaceutical industry due to its simplicity, accuracy, and versatility. Dr. Kitaguchi says, "Applying our screening method for the development of biopharmaceutical proteins may dramatically shorten the time required to establish highly productive industrial microbial strains. We thus believe that this study can contribute to the inexpensive manufacture of various biopharmaceutical proteins."
We hope these developments like these lead to a world where proteins can be produced more easily and affordably on demand!
Reference
Authors : |
Yoshihiro Ito1,2, Ryuichi Sasaki2, Sayaka Asari2, Takanobu Yasuda3, Hiroshi Ueda3, and Tetsuya Kitaguchi3 |
Title : |
Efficient Microfluidic Screening Method Using a Fluorescent Immunosensor for Recombinant Protein Secretions |
Journal : |
Small |
DOI : |
|
Affiliations : |
1Graduate School of Life Science and Technology, Tokyo Institute of Technology
2Research Institute for Bioscience Products and Fine Chemicals, Ajinomoto Co., Inc.
3Laboratory for Chemistry and Life Science, Institute of Innovative Research, Tokyo Institute of Technology
|
. Any information published on this site will be valid in relation to Science Tokyo.