A study led by researchers at Tokyo Institute of Technology (Tokyo Tech) has found that two key enzymes in plants called PAH1 and PAH21 are critical for survival and growth under nitrogen-depleted conditions. The study sheds new light on how plants could be modified in future to boost tolerance to nutrient-poor environments.
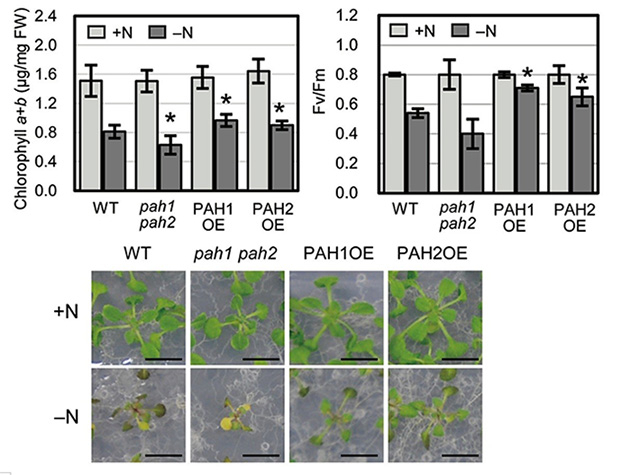
Figure 1. Comparison of chlorophyll content and photosynthetic activity under nitrogen starvation
Left: In nitrogen-depleted (–N, dark gray) conditions, plants overexpressing PAH1 and PAH2 (PAH1OE and PAH2OE) showed greater chlorophyll content than wild-type (WT) plants.
Right: PAH1OE and PAH2OE showed greater photosynthetic activity than WT plants under nitrogen-depleted (-N, dark gray) conditions.
Lower: Plants growth under normal growth (+N) and nitrogen depleted (-N) conditions. Scale bars: 1 cm
How plants tolerate nitrogen starvation is a longstanding mystery. Nitrogen is vital for the production of amino acids, the building blocks of plant proteins, and many other components needed to sustain life. Researchers in Japan have now found that two enzymes involved in lipid biosynthesis2 called PAH1 and PAH2 are essential for plant growth during nitrogen starvation. The finding advances fundamental knowledge of the processes regulating plant growth.
Published in Frontiers in Plant Science, the research was a result of collaboration between scientists from Tokyo Tech, the University of Tokyo and Tokyo University of Pharmacy and Life Sciences.
By studying a small flowering plant called Arabidopsis thaliana, the team showed that switching off two genes, PAH1 and PAH2, (in a process known as double knockout) led to increased sensitivity to nitrogen starvation. Arabidopsis is a popular choice among plant biologists due to its relatively short life cycle (of around two months) and small genome size (of around 135 megabase pairs), making it ideal for use as a model species.
The team compared the chlorophyll content and photosynthetic activity of the double knockout plants, transgenic plants that had been modified to produce more of (or overexpress) PAH1 and PAH2, and wild-type plants. The double knockout plants were found to have lower chlorophyll content than in the wild type under nitrogen-depleted conditions. Remarkably, the team found that transgenic plants showed a higher amount of chlorophyll and greater photosynthetic activity than the wild-type plants under nitrogen starvation (Figure 1).
Mie Shimojima of School of Life Science and Technology, Tokyo Tech, says that the study builds on around 20 years of work conducted by her research group on membrane lipid remodeling under inorganic phosphate (Pi)-depleted conditions.
"When plants suffer Pi starvation, phospholipids in the cellular membranes are degraded and replaced with glycolipids, or sugar-containing lipids; this is how plants survive the Pi shortage," says Shimojima. "In 2009, our colleagues Yuki Nakamura and others showed that PAH1 and PAH2 are crucial for plant growth under Pi-depleted conditions."
Growing evidence in recent years suggested that plant response to Pi starvation and nitrogen starvation might be related. "That's why we analyzed nitrogen starvation tolerance in the Arabidopsis plant lacking PAH1 and PAH2," says Shimojima. "Our study reinforces the view that the Pi starvation-induced lipid remodeling mechanism is also involved in the nitrogen starvation response."
"All of our findings so far indicate that PAH1 is involved in some kind of repair process or maintenance of chloroplast membrane structures," she continues. "However, since PAH is a cytosolic3 enzyme, there may be other essential proteins involved in this mechanism within the membrane."
Further studies will be needed to unravel the molecular mechanisms underlying nitrogen starvation tolerance and to explore how this knowledge may be used in agricultural and biotechnological applications.
1 PAH1 and PAH2
Phosphatidic acid phosphohydrolases — key regulatory enzymes.
2 Lipid biosynthesis
The reactions and pathways involved in the production of lipids, which provide energy for metabolic processes and structural components for cell membranes.
3 Cytosolic
Referring here to an enzyme found within the cytosol, the aqueous part of a plant cell.
Reference
Authors : |
Yushi Yoshitake1, Ryoichi Sato1, Yuka Madoka1, Keiko Ikeda1, Masato Murakawa1, Ko Suruga1, Daisuke Sugiura2, Ko Noguchi3, Hiroyuki Ohta1 and Mie Shimojima1,* |
Title of original paper : |
Arabidopsis Phosphatidic Acid Phosphohydrolases Are Essential for Growth under Nitrogen-Depleted Conditions |
Journal : |
Frontiers in Plant Science |
DOI : |
|
Affiliations : |
1Tokyo Institute of Technology, Japan
2The University of Tokyo, Japan
3Tokyo University of Pharmacy and Life Sciences, Japan
|
. Any information published on this site will be valid in relation to Science Tokyo.



