Genetic analyses conducted on an alga species may provide clues to the origin of auxin signaling mechanisms, as reported by scientists at Tokyo Tech. A series of experiments on Klebsormidium nitens — a member of an algae group sharing common ancestry with land plants unveiled key transcription factors and regulatory elements sensitive to auxin, an important hormone in land plants. The findings shed light on the diverging evolutionary paths of algae and land plants.
Phytohormones are chemical messengers that regulate the growth of plants and their response to the environment. In land plants, auxin is an important and well-studied phytohormone that affects various aspects of plant development. However, the main cell receptor that regulates the auxin signaling machinery in land plants—transport inhibitor response 1/auxin signaling F-box (TIR1/AFB)—is absent in algae. Past research has shown that a TIR1/AFB-independent auxin signaling mechanism may be involved in regulating the gene expression and growth in algae. However, not much is known about the underlying mechanisms of the auxin signaling pathway and their purpose in algae.
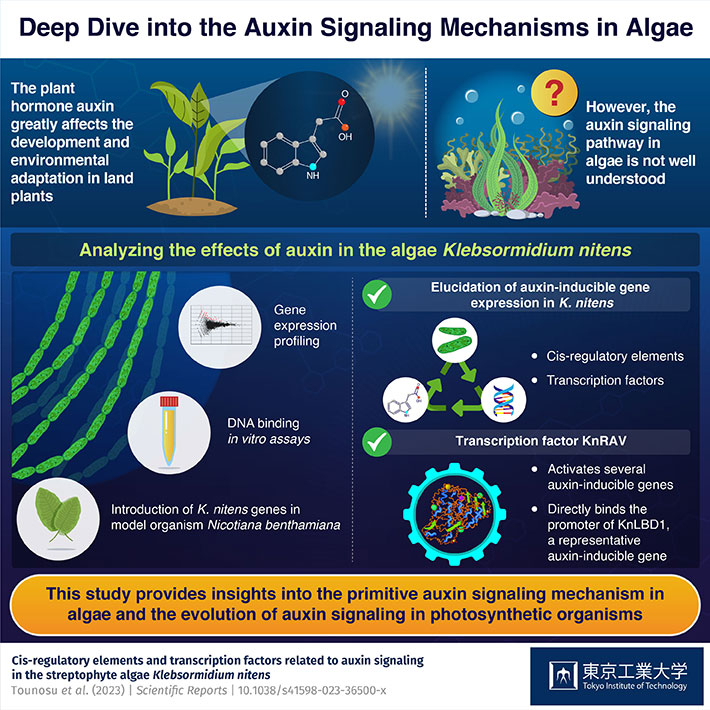
To address this knowledge gap, a team of scientists from the School of Life Science and Technology at Tokyo Institute of Technology (Tokyo Tech) in Japan conducted a study on the algae called Klebsormidium nitens. They selected this species because of two major reasons. Previous studies have shown that it produces indole-3-acetic acid (IAA), a type of auxin commonly seen in land plants. Also, external application of auxin halts cell proliferation in K. nitens, hinting that this species might have primitive auxin signaling mechanisms. The study was led by Assistant Professor Koichi Hori and was published in the journal Scientific Reports on 15 June 2023.
To this end, the team conducted a series of experiments on K. nitens, including gene expression profiling, RNA-sequence analysis, and DNA binding assays. In addition, they also introduced genes for various transcription factors potentially related to IAA signaling from K. nitens into Nicotiana benthamiana, a well-studied model organism in plant science with established analytical tools in scientific literature.
The experiments revealed that the RY motif was enriched in the promoter sequences of auxin-inducible genes in K. nitens. They also identified several cis-regulatory elements and transcription factors that reacted to the presence of IAA. Most importantly, they identified KnRAV — a transcription factor — as a key player in the auxin signaling mechanisms of K. nitens. "The results revealed that KnRAV activates several auxin-inducible genes and directly binds the promoter of KnLBD1, a representative auxin-inducible gene. Thus, KnRAV may have the potential to regulate auxin-responsive gene expression in K. nitens" explains Dr. Hori.
Overall, the researchers elucidate the intricacies of auxin signaling in algae via numerous experiments in K. nitens. Explaining the potential long-term important implications of these findings, Dr. Hori remarks, "The results provide new insights into the regulation of IAA-inducible gene expression in K. nitens. Moreover, the present findings not only enhance the understanding of auxin signaling in this alga, but also provide clues to discern the evolution of auxin signaling machinery in many other photosynthetic organisms."
We only wish that further studies can help elucidate the role of auxin in photosynthetic organisms!
. Any information published on this site will be valid in relation to Science Tokyo.