Transcript of Nobel Lecture by Yoshinori Ohsumi

on 7 December 2016
at the Aula Medica hall of the Karolinska Institutet
Ladies and gentlemen,
It is a great honor for me to be here as a recipient of the Nobel Prize in Physiology or Medicine. First I would like to express my sincere appreciation to the Karolinska Institute and Nobel foundation for this wonderful opportunity.
I would also like to thank everyone for attending today.
Let's begin.
Science is a system of knowledge that is gradually accumulated by society, but it is also an inherently human activity. I believe that every scientist is a product of the era in which they live. So today I'd like to start from a brief introduction of my life, and then give a historical overview of my scientific work.
I was born in Fukuoka, in Southern Japan, in 1945, half year before the end of World War II. This was a very challenging time in Japan, and everyone had difficulty getting basic daily necessities, including food. I myself suffered from severe malnutrition and was a very sickly child. Around this time my mother got tuberculosis and spent a long period bedbound, but she was miraculously able to recover thanks to a gift of just-developed antibiotics from a family friend in Hawaii. I remembered the words Streptomycin and Aureomycin, without fully understanding what they were.
My childhood home was surrounded by nature, with rice paddies, streams, hills and the sea all nearby. I spent a lot of time outdoors catching fish and picking plants. As an elementary school student, I was engrossed in collecting insects and watching the night's sky. I would say that this intimacy with nature had a strong influence on me. At the time, I vaguely longed to be a scientist.
In high school I joined the chemistry club, where I experienced the wonder of chemical reactions. I entered the University of Tokyo, where I initially hoped to become a chemist. But soon, I had difficulty finding a field to work in. Thankfully, this was the time when the central dogma was just being established. Molecular biology immediately fascinated me, so I decided to join the lab of Kazutomo Imahori as a graduate student. There I began my research career studying the ribosome, which is the machinery of protein synthesis. This period made me keenly aware of the continuous nature of protein synthesis within the cell. I gradually became interested in the uncharted function of bio-membranes, so I worked on the mechanism of action of colicin E3, which is a cytotoxic protein that inhibits protein synthesis by binding to a receptor on the membranes of other cells.
After I finished my doctoral studies, I enrolled in the lab of Gerald Edelman at The Rockefeller University, New York. During my final year in Dr. Edelman's lab I worked with Mike Jazwinski on the initiation mechanism of DNA replication in yeast, which was my first encounter with yeast as an experimental organism.
 Slide 3
Slide 3
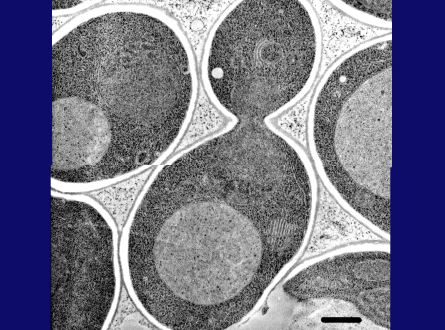
At the end of 1977 I returned to Japan as an assistant professor in Yasuhiro Anraku's lab at the University of Tokyo. Dr. Anraku's entire lab was tasked with studying the transporters and the respiratory chain in E. coli, but he fortunately allowed me to start yeast research. After a period of reflection, I decided to work on the yeast vacuole. I am not a competitive person, but rather prefer to work on a subject that is not fashionable. This EM picture shows a yeast vacuole, which is fairly large but has no obvious structures inside. At that time the vacuole was thought to be no more than a garbage dump in the cell, but I thought it must play yet unknown roles in cell physiology. I was able to show active transport of amino acids and calcium ions through the vacuolar membrane, providing evidence that the vacuole is important for homeostasis of metabolites and ions. We also described a novel proton-pump on the vacuolar membrane, the V-type ATPase, which generates a proton gradient across the vacuolar membrane.
In 1988 I was able to start my own lab — consisting of just myself — at the College of Arts and Sciences at the University of Tokyo. I decided to devote myself to the lytic function of the yeast vacuole. Since it is an acidic compartment and contains various hydrolytic enzymes, I thought it might be homologous to the lysosome. However, I had no concrete idea how to approach this problem.
Slide 7
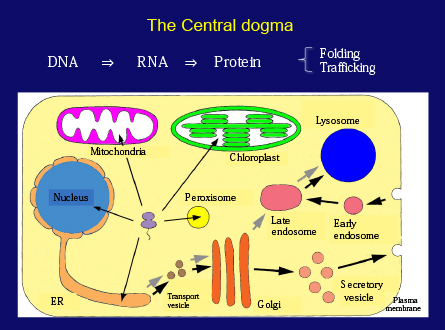
Before going into experimental details, I will briefly touch on protein metabolism in cells. Proteins, which are a polymer of amino acids, are the key players in all biological processes. The Central Dogma stated that protein is synthesized according to the genetic information encoded in DNA through RNA. Since then, many researchers have studied hard to understand gene expression and protein synthesis. Recent progress in cell biology has revealed how each protein localizes to its specific destination in the cell. We now know that each protein is ingeniously organized through the processes of intracellular trafficking.
However, one aspect that was missing from this picture was the degradation of proteins within cells. When I held a biology class for first-year undergraduate students, I used to start my lecture with this question: how many red blood cells are made within just one second in your body? Calculation is quite simple and gives an answer of about three million cells per second. How about hemoglobin in the red blood cell? Further calculation shows that about 1×1015, or one million billion, hemoglobin molecules are synthesized per second. It follows that exactly the same amount of cells and proteins are degraded. The key message is that living things are so dynamically maintained.
The Japanese climate is characterized by four distinct seasons that influence our culture, and students at school learn that all things are continuously changing. Plant leaves offer an excellent example. In spring, leaves develop and then actively synthesize starch by sunlight, but in autumn leaves turn yellow and fall off. What we see as beautiful autumn colors is actually caused by the degradation of the leaf's photosynthetic machinery: green chloroplasts are completely degraded, and the resulting amino acids are transported to the trunk. Similarly rice leaves turn yellow at harvest-time. All the protein in the leaves is degraded and transported to make proteins in rice grains for the next generation. These examples demonstrate that degradation is not an adverse process, but rather is essential for new construction or regeneration. Let's think about the human body. Our bodies make about 2-300 grams of protein every day, but we only obtain about 70-80 grams of protein from our diet. The amino acids required for protein synthesis mostly come from the degradation of the body's own proteins. Life is maintained by a tightly controlled balance between synthesis and degradation, and the recycling of protein is essential for life.
The earliest pioneer in the field of protein metabolism was Rudolph Shoenheimer, who, in the 1930s, proposed the concept of protein turnover. Unfortunately his idea was not widely accepted. A monumental breakthrough came in the 1950s with the discovery of the lysosome by Christian de Duve. Soon after, researchers at Rockefeller University analyzed the process of delivery of own cellular components to lysosomes by electron microscopy, a process which de Duve coined "autophagy" in 1962, from the Greek for "self-eating". Glen Mortimore and Per Seglen afterwards produced informative works by tackling both the physiological meaning and regulation of autophagy on mammals. Despite this, our understanding of the molecular mechanisms of autophagy remained in its infancy for many years.
Slide 17
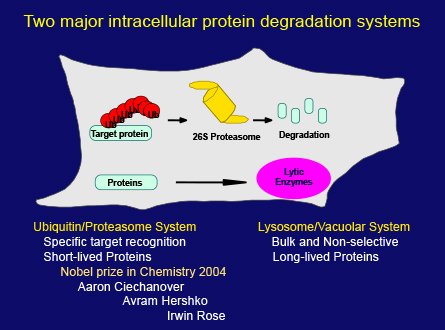
In the meantime, another intracellular degradation system, the ubiquitin/proteasome system, was discovered. As it became clear that this system plays important roles in the regulation of a variety of cellular processes, protein degradation suddenly attracted a huge amount of attention in biological research. This was ultimately recognized by the Nobel Prize in Chemistry in 2004 which was awarded to these three scientists shown here, but the field of lysosomal degradation and autophagy still remained quiet.
Now let's return to my work. Assuming that the vacuole functions as a lytic compartment, the questions I had were when, what and how cytoplasmic constituents go across the vacuolar membrane and become accessible to vacuolar enzymes. First I searched for a condition where massive protein degradation might occur in the yeast life cycle. Spore formation is a meiotic process that generates four spores and is induced by the depletion of nitrogen, the essential element for amino acids. I reasoned that this dramatic cellular remodeling should require the degradation of pre-existing proteins to make necessary proteins.
I have to admit that I've always loved to observe yeast cells by light microscopy. The vacuole is relatively large and is the only clearly visible organelle in the cell, and the inside of the vacuole is just like a salt solution. So I knew that it is easy to see structures if they are inside the vacuole. I probably spent more time sitting at the light microscope than anyone else at that time.
I carefully observed the early stages of the sporulation process, but couldn't see any significant changes in the cell. I thought that this may be due to the degradation of structures in the vacuole. I conceived to use a vacuolar proteinase deficient mutant to prevent further degradation. Fortunately, Elizabeth Jones had donated many proteinase mutants to the Yeast Genetic Stock Center, so I obtained these strains and shifted to nitrogen depleted medium.
Looking down the microscope, I saw many spherical structures vigorously moving around in the vacuole. These structures appeared within 30 minutes and almost completely filled the vacuole after 3-4 hours. For me, this observation marked the exact starting point of more than 27 years of autophagy research. The reason that I could discover this phenomenon with only a common low magnification light microscope owes to the vigorous movement of these structures, by Brownian motion, in the otherwise static yeast cell. This dramatic change was enough to convince me that I had encountered an unknown and fascinating phenomenon. Soon Kazuhiko Takeshige showed that this phenomenon is not restricted to sporulation, but is a general cellular response to a range of nutrient starvation conditions.
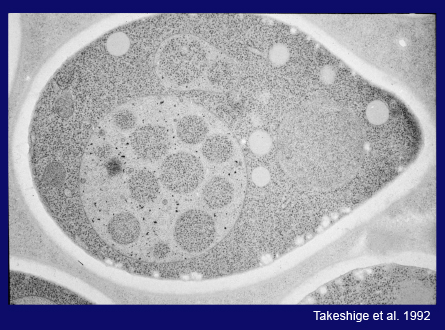
I then called on the excellent electron microscopists Misuzu Baba and Masako Osumi to capture the morphology of this phenomenon within cells by EM. Their efforts produced beautiful and conclusive images of the entire autophagy process, as shown in the following several slides.
Slide 24
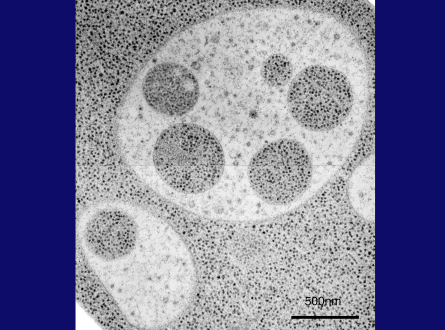
Slide 25
This is a thin section image of a nitrogen starved cell at 3 hrs. You can see spherical structures within vacuoles, and this high magnification image shows these structures are bound by a single membrane and their contents are exactly the same as a portion of cytoplasm, containing the same density of ribosomes and various cytoplasmic structures, such as the rough ER here, suggesting nonselective sequestration of cytoplasmic components. We also observed a membrane sac, the isolation membrane, just enwrapping a portion of cytoplasm next to the vacuole. After completion of membrane expansion and sealing, a double membrane structure, the yeast autophagosome, is formed. This freeze-fracture image shows the fusion event of an autophagosome with the vacuolar membrane, where the outer membrane of the autophagosome is continuous with the vacuolar membrane. Here we can see inner membrane structures, which we named autophagic bodies. Autophagic bodies occasionally contain mitochondria, as you can see here and here, indicating that autophagy can degrade not only cytosolic protein but also supramolecular structures such as ribosomes, and even entire organelles, which is a characteristic feature of autophagy.
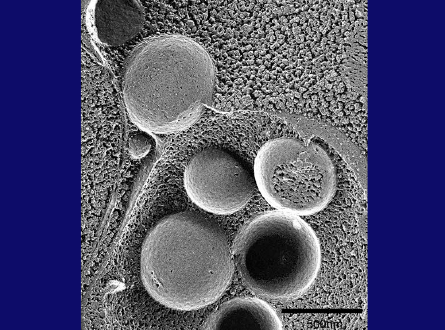
Slide 28
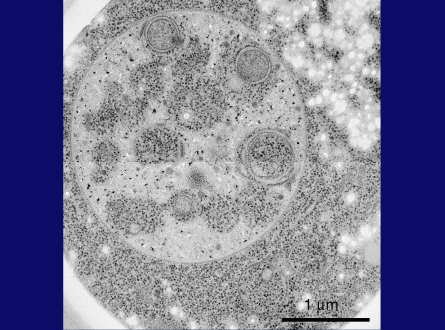
Slide 29
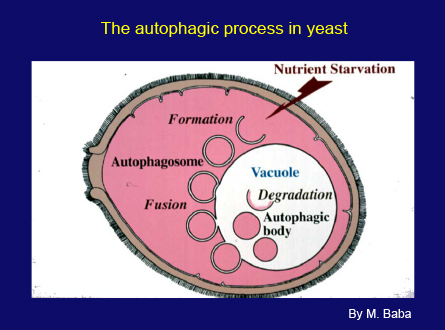 Slide 30
Slide 30
This slide shows a scheme of the autophagic process in yeast. When cells are faced with starvation conditions, a small membrane sac appears next to the vacuole and expands to enwrap a portion of cytoplasm. Then, a double membrane structure, the autophagosome, targets to the vacuole and fuses with the vacuolar membrane, releasing the inner membrane structure into the vacuole. In wild type cells these structures are immediately disrupted by vacuolar enzymes. The proteinase-deficient mutant allowed us to follow the progression of autophagy as the accumulation of autophagic bodies. The membrane dynamics of this process are essentially the same as known mammalian autophagy, although the vacuole is much larger than the lysosome.
As you know, yeast is a very good organism for the dissection of complicated biological phenomena through genetic analyses. I was inspired by the beautiful works of Lee Hartwell, which unlocked the cell cycle through cdc mutants, and also by the uncovering of the secretory pathway using sec mutants by Randy Schekman.
We therefore set out to obtain autophagy-defective mutants to find the genes involved in autophagy. However, at the time we had no idea how an autophagy-defective mutant would behave, so we decided to perform a simple morphological screen for mutants that don't accumulate autophagic bodies under starvation conditions. Miki Tsukada, one of my first graduate students, made a great contribution during this screening process. By assessing thousands of mutagenized cells one by one for mutants lacking autophagic bodies, she identified the first autophagy-defective mutant, which we named apg1-1, later renamed to Atg1. This mutant obviously failed to degrade proteins upon starvation. But it grew normally in rich medium, and otherwise had no obvious phenotype. However, we soon found that Atg1 mutants die sooner than wild type cells during long periods of starvation. Assuming that this phenotype is caused by a defect in autophagy, Miki ran another screen assessing the viability of mutagenized cells first and successfully obtained about one hundred autophagy-defective mutants. Genetic analysis of these strains revealed 14 Atg mutants. We now know that 18 Atg genes are essential for starvation-induced autophagy in yeast, so I can say that this initial screen was very effective. Soon, our work using electron microscopy suggested that all these Atg mutants are defective in the formation of the autophagosome, the most crucial event in autophagy.
Obviously, the next step was to determine the proteins encoded by each Atg gene. The first gene, Atg1, turned out to encode a protein kinase. But most of the Atg genes cloned by our group and others were completely novel, and their amino acid sequences did not tell us anything about their functions. This was a difficult time for us.
My lab in the College of Arts and Sciences was quite small and had no fancy equipment. Funding was limited, and I had only a few colleagues. But it is without doubt that the eight years in that lab are the foundation of my work in autophagy.
In 1996, I became a professor at the National Institute of Basic Biology in Okazaki, which provided us with a very conducive research environment. I asked Tamotsu Yoshimori to join our lab as an associate professor to start work on autophagy in mammalian cells, and also Takeshi Noda and Yoshi Kamada to join us as assistant professors. The following year, Noboru Mizushima came to my lab as a Post Doc. Year by year, talented postdoctoral researchers and graduate students gathered to my lab. The cloning of Atg genes proceeded much faster than I expected thanks to the collaboration of Mariko Ohsumi's lab and the completion of the yeast whole genome project.
The first breakthrough was made by Noboru Mizushima. When he examined the product of Atg12 he found two bands, one of which ran at a much higher apparent molecular weight. He also realized that in Atg5, Atg7, Atg10 mutants this high molecular weight band was missing. Although I won't go into details, we found that Atg12, a novel ubiquitin-like molecule is activated by the E1 enzyme Atg7 and then transferred to the E2 enzymeAtg10, finally formed a conjugate with a sole target protein, Atg5. We also found that two Atg12-Atg5 conjugates were combined by an Atg16 dimer. The discovery of this enzyme system uncovered the function of 5Atg proteins all at once.
Another molecule we were interested in was Atg8, because immunoelectron microscopy showed that Atg8 is localized to autophagy-related membranes, suggesting that it is a good marker for membrane biogenesis. Two talented graduate students, Takayoshi Kirisako and Yoshinobu Ichimura, unveiled another unique conjugation system. Atg8 is synthesized as a precursor that is processed by the proteinase Atg4 and is then activated by the same E1 enzyme, Atg7, and then transferred to Atg3. But rather than forming a conjugate with a protein, it forms a bond with one of the major membrane phospholipids, phosphatidylethanolamine. Thus to our surprise we found that about half of the Atg genes are involved in these two conjugation systems.
In 1998 Takeshi Noda found that rapamycin induces autophagy even when cells are grown in rich medium. As rapamycin is an inhibitor of TOR kinase, we therefore proposed that TOR kinase is the most upstream regulator of autophagy. Meanwhile, Yoshi Kamada and Tomoko Funakoshi studied the early steps of autophagy and showed that Atg13 and Atg17 play an important role in autophagy induction and are necessary for Atg1 kinase activation. Akio Kihara also found that autophagy requires the specific PI3 kinase complex I, which is distinct from the predominant Complex II.
Within a decade of our research at NIBB, we found two more Atg proteins, which interact with each other as well as Atg17 and regulate Atg1 kinase activity. We concluded that a total of 18 Atg proteins are essential for autophagy, and that these proteins can be classified into 6 functional groups: the Atg1 kinase and its regulators, the specific PI3 kinase complex, two conjugation systems, the sole membrane spanning membrane protein Atg9, and the Atg2-Atg18 complex.
Also during our time at NIBB, Noboru Mizushima and Tamotsu Yoshimori began to study mammalian autophagy, and successfully showed that the Atg12 conjugation system is well conserved in mammalian cells. Tamotsu definitively proved that LC3, the yeast Atg8 homologue, can be used as a very effective marker of autophagy progression. At around this time, two of my colleagues, Hideki Hanaoka and Kohki Yoshimoto, succeeded in identifying Atg genes in plant cells. Although the homology of Atg genes is low, the Atg system is well conserved between yeast, mammals and plants, indicating that the core machinery of autophagy was acquired at an early stage of evolution.
The identification of Atg genes essential for autophagy changed the entire landscape of autophagy research. One aspect of this revolution is that we can now visualize the process of autophagy by fluorescence microscopy using Atg proteins as markers. Noboru Mizushima took advantage of this when he constructed a transgenic mouse expressing GFP-LC3. This tool has allowed us to assess how much autophagy occurs in every organ of the mouse's body. This mouse has been shared with labs all over the world. Noboru also developed the first Atg gene knockout mice, which allowed us to show that autophagy plays a crucial role for survival at birth. Masaki Komatsu was also able to demonstrate that conditional knockout of the Atg7 gene causes accumulation of ubiquitinated proteins and ultimately tumor in the liver. From this point, manipulation of Atg genes in various organisms, different cell types, a variety of tissues and organs, and individuals started in many labs around the world and revealed a vast array of physiological functions of autophagy. The relevance of autophagy in various diseases is becoming clear. This rapid expansion of autophagy would not have been possible without the concerted efforts of autophagy researchers all over the world. But this huge area falls outside the scope of my talk today, which focuses on my fundamental research in yeast.
I mentioned earlier that nutrient recycling is the most primary function of autophagy, which is an evolutionary adaptation to the persistent challenge of nutrient limitation, starvation. Recent works have shown another important function of autophagy. In addition to the bulk recycling of portions of cytoplasm, autophagy is also able to pick and choose excess or harmful materials from the cytoplasm in a selective manner. Targets of this degradation include specific proteins, supramolecular complexes, protein aggregates, organelles and even viruses and invasive bacteria. Now it is well accepted that autophagy plays an essential role in the maintenance of cytoplasmic homeostasis. Defects in this quality control may cause various abnormalities, health problems or diseases. This kind of autophagy is generally not random, but selective. Also using yeast, Dan Klionsky has been studying the Cvt pathway, the first identified model system of selective autophagy. In yeast several receptors for selective autophagy have been identified. Now the various selective modes of autophagy have become one of the most exciting fields in autophagy research. We are only now beginning to arrive at a clear understanding for some of these selective types of autophagy, such as mitochondrial autophagy, but this is an area in which we need further research to obtain the precise molecular details.
Slide 52
Slide 63
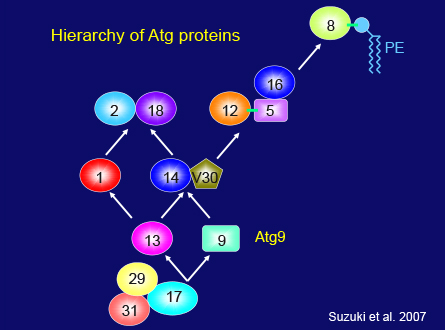
Even in the simple yeast system many fundamental problems remain to be answered. For this reason my group is still working on yeast autophagy. Although we had identified the core components in autophagosome formation, the questions of where and how these proteins function in the cell remained. Kuninori Suzuki made a major contribution in this area. First he found that when autophagy is induced most Atg proteins localize at least partly to a dot structure next to the vacuole. He also showed that the autophagosome is generated from this dot structure, which we called the preautophagosomal structure, or the PAS. We systematically analyzed the mutual relationship between Atg proteins to uncover how the PAS is organized, and found a causal relationship among the 6 functional groups, as shown in this slide, which gave us a framework for further studies. Recent developments in fluorescence microscopy have allowed us to observe Atg proteins in the cell, showing that most Atg proteins move freely in the cytoplasm under growing conditions. Upon starvation, only a small portion of each Atg protein is recruited to the PAS. We now know that the hierarchical relationship between Atg proteins reflects the temporal sequence of events that occur during autophagosome formation.
Close and long term collaboration with Nobuo Noda and the late Dr. Fuyuhiko Inagaki successfully provided us with structural information of many of the Atg proteins and their complexes, which was really crucial for our understanding of their function.
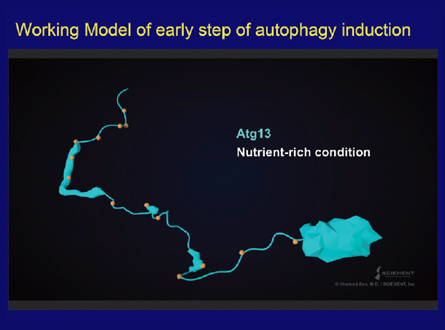
Now I would like to talk briefly about the recent works on the early steps of autophagosome formation, which were mostly done by Hayashi Yamamoto and Nobuo Noda. Atg13, a key player, has a unique structure that is made up of a N-terminal globular domain and a long naturally disordered region. This disordered region contains many phosphorylated amino acids by TOR1 kinase. Upon starvation Atg13 is quickly dephosphorylated, which allows it to bind to Atg1 and Atg17 via specific sites in the disordered region, resulting in the formation of a dimer of the Atg1, Atg13, Atg17, Atg29 and Atg31 complexes. Work by Dan Klionsky's group had suggested that the actual number of Atg proteins assembling at the PAS is likely to be 30-40. How is this further assembly realized? We found that Atg13 has a second, weak Atg17 binding site that is also essential for PAS formation and autophagy. Now I will present our image of the early organization of the PAS in this movie. As you can see, Atg13 has an N-terminal globular domain and long naturally disordered region. In this region of Atg13, Atg1 kinase binds as shown. Then, the Atg17-29-31 complex is recruited via an Atg17 binding site. We found that the second biding site, shown here, is essential for PAS formation and autophagy. Atg13 is not able to use both of the binding sites in a single Atg17 protein, but rather links two Atg17 molecules, resulting in the PAS scaffold. By this way, this scaffold can form a large assembly. This assembly is essential for the activation of Atg1 kinase via auto-phosphorylation between adjacent Atg1 molecules, which is necessary for the next step of membrane formation. Recently, Sho Suzuki found that the N-terminal domain of Atg13 binds to the N-terminal disordered region of Atg9, which allows the first step of the recruitment of Atg9 vesicles to the PAS. These findings together demonstrate that the PAS is not static, but is in fact a dynamic and flexible structure consisting of a large number of Atg proteins and membrane structures. During membrane formation, the PAS must be regulated spatiotemporally in a concerted manner by protein modification and the formation of a transient assembly. We still need to extend our analyses to the later steps of autophagosome formation.
At present my group is focused on the physiological meaning of autophagy in yeast. In a way, I feel that we have come full circle by going back to the original questions — when, how, and what is degraded by autophagy? By using the latest analytical tools, Hunag Huang and Tomoko Kawamata showed massive RNA degradation by autophagy and that the resulting bases are secreted out from the cell. We need to know more about the degradation process and assess the degradation products of autophagy, and how they are transported out of the vacuole to the cytoplasm, and how they affect cellular metabolism.
 Slide 67
Slide 67
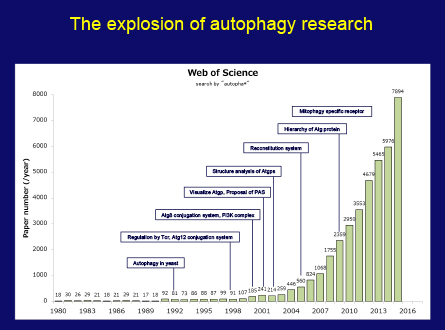
My autophagy research has always been driven by nothing more than intellectual curiosity and a thirst to get a better understanding of life through protein dynamics of the cell. When I started my work, I never thought it would become relevant in diseases as diverse as neuro-degeneration, infectious disease, cancer and others in such a short time. But now autophagy research has become a major field in biology. This figure presents that the number of papers related to autophagy is so rapidly increasing. It goes without saying that this is thanks in large part to the tremendous efforts of many researchers all over the world, so I would like to express my sincere thanks and share this honor with all of them.
This figure also tells us it takes a while to establish one field. Truly original discoveries in science are often triggered by unpredictable and unforeseen small findings. Nowadays, the distance between basic discoveries and practical applications is getting closer. While this is exciting, scientists are increasingly required to provide evidence of immediate and tangible applications of their work. It is my sincere hope that society is able to nurture not only purpose-oriented science, but also science as a core cultural activity.
If my small idea and decades of work have made a contribution to fundamental science through autophagy, and the Nobel Foundation recognizes the basic nature of this work with this prize, it is honestly my greatest pleasure and satisfaction as a basic scientist.
Finally I must acknowledge the good fortune I've had throughout my career, the excellent colleagues who have joined me on this journey, the great efforts of many collaborators, good friends, the continuous support of many grants and my caring family, especially my wife Mariko.
Thank you for your attention.
* For more information about the slides of the Nobel lecture, please visit following website
Published: July 2017
. Any information published on this site will be valid in relation to Science Tokyo.













